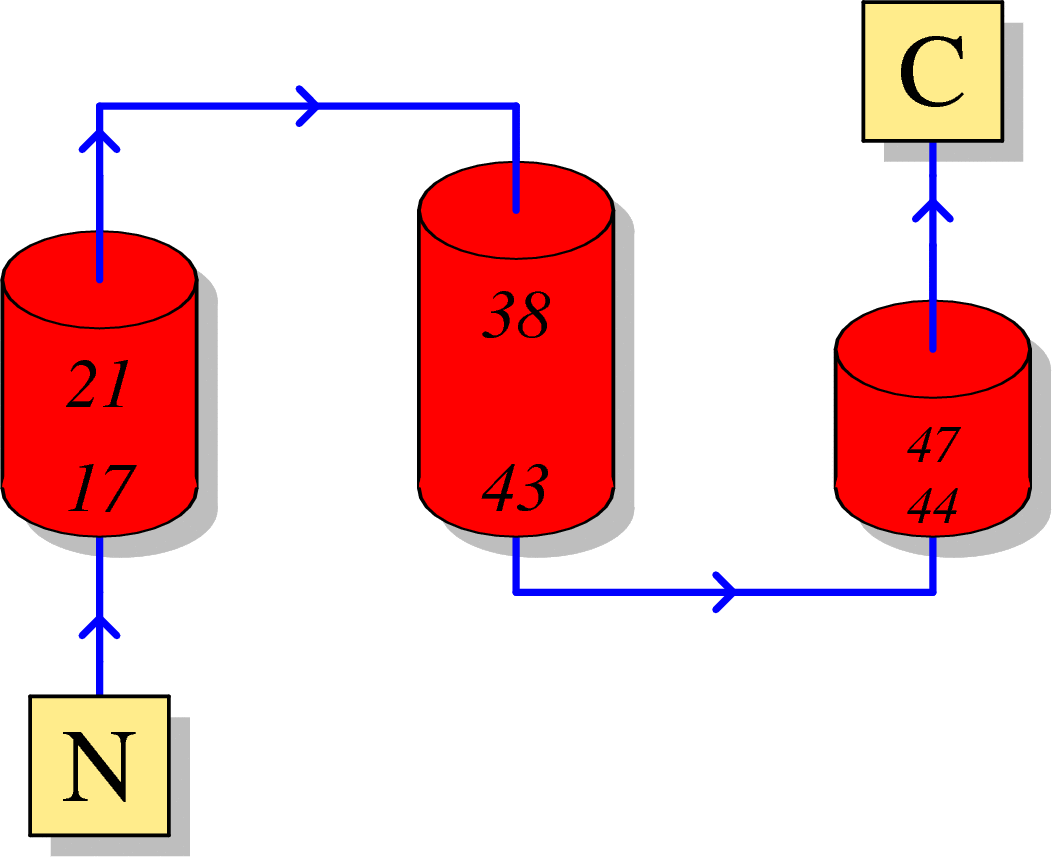
Function of HIV p6
p6 is a structural protein at the C terminus of gag gene [1].
p6 can recruit the host machinery to bud the virus outwards from the cell surface [2].
Viral protein Vpr and host proteins (e.g. AIP1/ALIX) bind to p6 during viral packaging [1].
Reference
Bell NM, Lever AM: HIV Gag polyprotein: processing and early viral particle assembly. Trends Microbiol 2013, 21:136-144.(Download Article)
Morita E, Sundquist WI: Retrovirus budding. Annu Rev Cell Dev Biol 2004, 20:395-425.(Download Article)
Sequence
(1) Reference sequence for HIV-1 p6
1 10 20 30 40 50
| | | | | |
LQSRPEPTAP PEESFRSGVE TTTPPQKQEP IDKELYPLTS LRSLFGNDPS
51
|
SQ (2) Reference sequence for HIV-2 and SIV p6
1 10 20 30 40 50
| | | | | |
PMAQVHQGLM PTAPPEDPAV DLLKNYMQLG KQQREKQRES REKPYKEVTE
51 60
| |
DLLHLNSLFG GDQ* (3) Coloring scheme for above amino acids
Amino acids with hydrophobic side chains (normally buried inside the protein core):
A - Ala - Alanine
I - Ile - Isoleucine
L - Leu - Leucine
M - Met - Methionine
V - Val - Valine
Amino acids with polar uncharged side chains (may participate in hydrogen bonds):
N - Asn - Asparagine
Q - Gln - Glutamine
S - Ser - Serine
T - Thr - Threonine
Amino acids with positive charged side chains:
H - His - Histidine
K - Lys - Lysine
R - Arg - Arginine
Amino acids with negative charged side chains:
D - Asp - Aspartic acid
E - Glu - Glutamic acid
Amino acids with aromatic side chains:
F - Phe - Phenylalanine
Y - Tyr - Tyrosine
W - Trp - Tryptophan
Cysteine: C - Cys - Cysteine
Glycine: G - Gly - Glycine
Proline: P - Pro - Proline
Amino acid variations at HIV-1 p6
Here, we visualize the prevalence of amino acid variations at the HIV-1 p6 from HIV-1 subtype B.
Protocal of our sequence collection
For HIV-1 subtype B, one sequence per patient was extracted from HIV Los Alamos database (www.hiv.lanl.gov/).
We removed misclassified sequences or sequences with hypermutations, stop codons, ambiguous nucleotides, which were described in our article [1].
We removed sequences conferred partial or full resistance to any of the protease inhibitors, RT inhibitors and integrase inhibitors using HIVdb V6.0 .
Visualization
Our sequence dataset of HIV-1 subtype B p6 included 4725 sequences. In the following picture, HXB2 indices of individual proteins are shown on top of the colored bars. A consensus amino acid at each position is shown beneath the colored bar. Natural variations are shown below the consensus amino acids; proportions (%) are colored red if they were more than 5%; blue otherwise.
HIV-1 protein interaction patterns.
Please cite our article:
Guangdi Li, Supinya Piampongsant, Nuno Rodrigues Faria, Arnout Voet, Andrea-Clemencia Pineda-Peña, Ricardo Khouri, Philippe Lemey, Anne-Mieke Vandamme, Kristof Theys. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18, doi:10.1186/s12977-015-0148-6 (2015). [PDF] [PubMed Link]