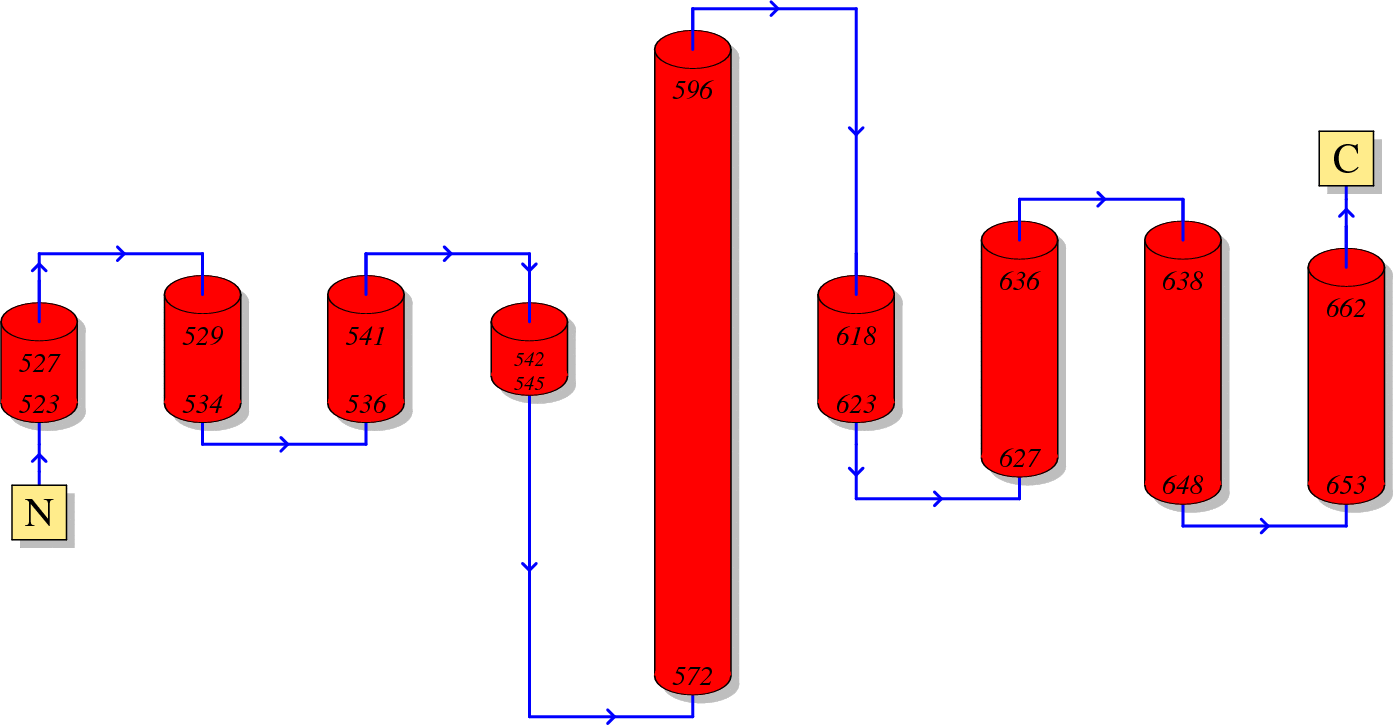
Function of HIV GP41
Encoded by the env gene, the surface glycoprotein GP41 is present on the surface of HIV particles.
GP41 contains a glycine-rich region which is essential for the membrane fusion activity [1].
The intracellular trafficking of the Env protein is regulated by the cytoplasmic tail of GP41 (GP41CT) which interacts with various cellular proteins.
GP41CT interacts with the viral Matrix protein to regulate the Env incorporation into HIV virions.
GP41CT regulates the internalization exerted by the clathrin-mediated endocytosis.
GP41CT regulates cellular activation of host transcription factors (e.g. NF-kB).
GP41 interacts with host proteins to regulate the activity of actin cytoskeleton.
HIV-1 GP41 membrane-proximal external region is targeted by human antibodies (e.g. 10E8) with a broad neutralization activity [2].
Reference
Postler TS, Desrosiers RC: The tale of the long tail: the cytoplasmic domain of HIV-1 gp41. Journal of virology 2013, 87(1):2-15.(Download Article)
Huang J, Ofek G, Laub L, Louder MK, Doria-Rose NA, Longo NS, Imamichi H, Bailer RT, Chakrabarti B, Sharma SK et al: Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 2012, 491(7424):406-412.(Download Article)
Sequence
(1) Reference sequence for HIV-1 GP41
1 10 20 30 40 50
| | | | | |
AVGIGALFLG FLGAAGSTMG AASMTLTVQA RQLLSGIVQQ QNNLLRAIEA
51 60 70 80 90 100
| | | | | |
QQHLLQLTVW GIKQLQARIL AVERYLKDQQ LLGIWGCSGK LICTTAVPWN
101 110 120 130 140 150
| | | | | |
ASWSNKSLEQ IWNHTTWMEW DREINNYTSL IHSLIEESQN QQEKNEQELL
151 160 170 180 190 200
| | | | | |
ELDKWASLWN WFNITNWLWY IKLFIMIVGG LVGLRIVFAV LSIVNRVRQG
201 210 220 230 240 250
| | | | | |
YSPLSFQTHL PTPRGPDRPE GIEEEGGERD RDRSIRLVNG SLALIWDDLR
251 260 270 280 290 300
| | | | | |
SLCLFSYHRL RDLLLIVTRI VELLGRRGWE ALKYWWNLLQ YWSQELKNSA
301 310 320 330 340 345
| | | | | |
VSLLNATAIA VAEGTDRVIE VVQGACRAIR HIPRRIRQGL ERILL (2) Reference sequence for HIV-2 and SIV GP41
1 10 20 30 40 50
| | | | | |
GVFVLGFLGF LATAGSAMGA ASLTLTAQSR TLLAGIVQQQ QQLLDVVKRQ
51 60 70 80 90 100
| | | | | |
QELLRLTVWG TKNLQTRVTA IEKYLKDQAQ LNAWGCAFRQ VCHTTVPWPN
101 110 120 130 140 150
| | | | | |
ASLTPKWNNE TWQEWERKVD FLEENITALL EEAQIQQEKN MYELQKLNSW
151 160 170 180 190 200
| | | | | |
DVFGNWFDLA SWIKYIQYGV YIVVGVILLR IVIYIVQMLA KLRQGYRPVF
201 210 220 230 240 250
| | | | | |
SSPPSYFQQT HIQQDPALPT REGKERDGGE GGGNSSWPWQ IEYIHFLIRQ
251 260 270 280 290 300
| | | | | |
LIRLLTWLFS NCRTLLSRVY QILQPILQRL SATLQRIREV LRTELTYLQY
301 310 320 330 340 350
| | | | | |
GWSYFHEAVQ AVWRSATETL AGAWGDLWET LRRGGRWILA IPRRIRQGLE
351
|
LTLL (3) Coloring scheme for above amino acids
Amino acids with hydrophobic side chains (normally buried inside the protein core):
A - Ala - Alanine
I - Ile - Isoleucine
L - Leu - Leucine
M - Met - Methionine
V - Val - Valine
Amino acids with polar uncharged side chains (may participate in hydrogen bonds):
N - Asn - Asparagine
Q - Gln - Glutamine
S - Ser - Serine
T - Thr - Threonine
Amino acids with positive charged side chains:
H - His - Histidine
K - Lys - Lysine
R - Arg - Arginine
Amino acids with negative charged side chains:
D - Asp - Aspartic acid
E - Glu - Glutamic acid
Amino acids with aromatic side chains:
F - Phe - Phenylalanine
Y - Tyr - Tyrosine
W - Trp - Tryptophan
Cysteine: C - Cys - Cysteine
Glycine: G - Gly - Glycine
Proline: P - Pro - Proline
Amino acid variations at HIV-1 GP41
Here, we visualize the prevalence of amino acid variations at the HIV-1 GP41 from HIV-1 subtype B.
Protocal of our sequence collection
For HIV-1 subtype B, one sequence per patient was extracted from HIV Los Alamos database (www.hiv.lanl.gov/).
We removed misclassified sequences or sequences with hypermutations, stop codons, ambiguous nucleotides, which were described in our article [1].
We removed sequences conferred partial or full resistance to any of the GP41 inhibitors, RT inhibitors and GP41 inhibitors using HIVdb V6.0 .
Visualization
Our sequence dataset of HIV-1 subtype B GP41 included 4725 sequences. In the following picture, HXB2 indices of individual proteins are shown on top of the colored bars. A consensus amino acid at each position is shown beneath the colored bar. Natural variations are shown below the consensus amino acids; proportions (%) are colored red if they were more than 5%; blue otherwise.
HIV-1 protein interaction patterns.
Please cite our article:
Guangdi Li, Supinya Piampongsant, Nuno Rodrigues Faria, Arnout Voet, Andrea-Clemencia Pineda-Peña, Ricardo Khouri, Philippe Lemey, Anne-Mieke Vandamme, Kristof Theys. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18, doi:10.1186/s12977-015-0148-6 (2015). [PDF] [PubMed Link]