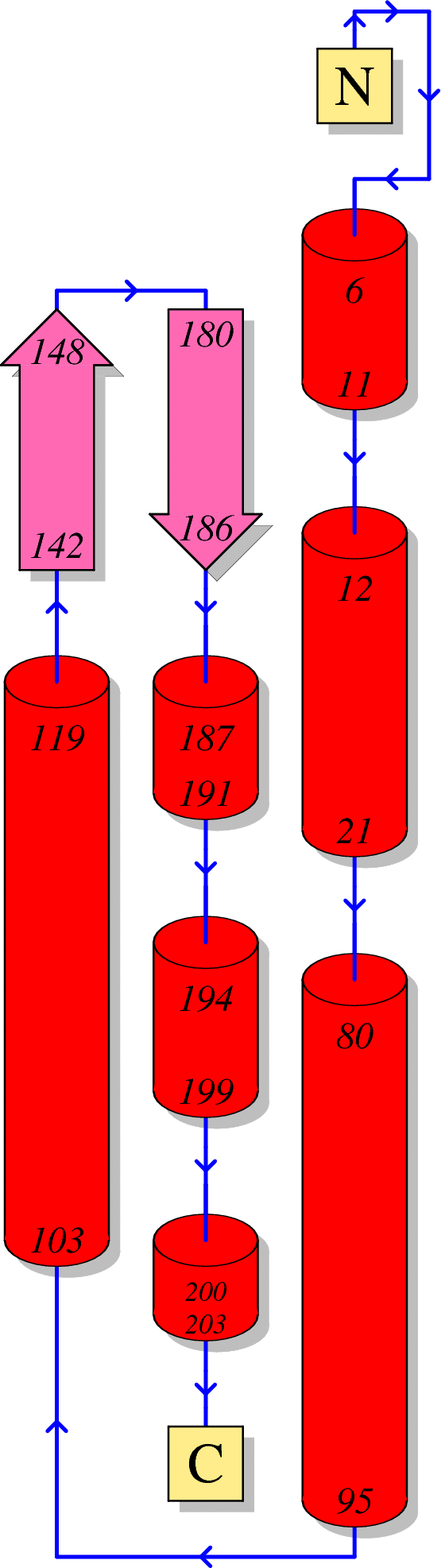
Function of HIV Nef
Negative regulatory factor (Nef) is an accessory protein which enhances viral pathogenesis [1].
Nef downregulates CD4 receptors and MHC molecules [1].
Nef promotes the viral release and the cell-to-cell transmission.
Nef activates the apoptosis and involves with the clathrin-dependent endocytic pathways. Nef is incorporated in the viral particles [1].
Reference
Laguette N, Bregnard C, Benichou S, Basmaciogullari S: Human immunodeficiency virus (HIV) type-1, HIV-2 and simian immunodeficiency virus Nef proteins. Molecular aspects of medicine 2010, 31(5):418-433.(Download Article)
Sequence
(1) Reference sequence for HIV-1 Nef
1 10 20 30 40 50
| | | | | |
MGGKWSKSSV IGWPTVRERM RRAEPAADRV GAASRDLEKH GAITSSNTAA
51 60 70 80 90 100
| | | | | |
TNAACAWLEA QEEEEVGFPV TPQVPLRPMT YKAAVDLSHF LKEKGGLEGL
101 110 120 130 140 150
| | | | | |
IHSQRRQDIL DLWIYHTQGY FPD*QNYTPG PGVRYPLTFG WCYKLVPVEP
151 160 170 180 190 200
| | | | | |
DKIEEANKGE NTSLLHPVSL HGMDDPEREV LEWRFDSRLA FHHVARELHP
201 206
| |
EYFKNC (2) Reference sequence for HIV-2 and SIV Nef
1 10 20 30 40 50
| | | | | |
MGGAISMRRS RPSGDLRQRL LRARGETYGR LLGEVEDGYS QSPGGLDKGL
51 60 70 80 90 100
| | | | | |
SSLSCEGQKY NQGQYMNTPW RNPAEEREKL AYRKQNMDDI DEEDDDLVGV
101 110 120 130 140 150
| | | | | |
SVRPKVPLRT MSYKLAIDMS HFIKEKGGLE GIYYSARRHR ILDIYLEKEE
151 160 170 180 190 200
| | | | | |
GIIPDWQDYT SGPGIRYPKT FGWLWKLVPV NVSDEAQEDE EHYLMHPAQT
201 210 220 230 240 250
| | | | | |
SQWDDPWGEV LAWKFDPTLA YTYEAYVRYP EEFGSKSGLS EEEVRRRLTA
251 260
| |
RGLLNMADKK ETR (3) Coloring scheme for above amino acids
Amino acids with hydrophobic side chains (normally buried inside the protein core):
A - Ala - Alanine
I - Ile - Isoleucine
L - Leu - Leucine
M - Met - Methionine
V - Val - Valine
Amino acids with polar uncharged side chains (may participate in hydrogen bonds):
N - Asn - Asparagine
Q - Gln - Glutamine
S - Ser - Serine
T - Thr - Threonine
Amino acids with positive charged side chains:
H - His - Histidine
K - Lys - Lysine
R - Arg - Arginine
Amino acids with negative charged side chains:
D - Asp - Aspartic acid
E - Glu - Glutamic acid
Amino acids with aromatic side chains:
F - Phe - Phenylalanine
Y - Tyr - Tyrosine
W - Trp - Tryptophan
Cysteine: C - Cys - Cysteine
Glycine: G - Gly - Glycine
Proline: P - Pro - Proline
Amino acid variations at HIV-1 Nef
Here, we visualize the prevalence of amino acid variations at the HIV-1 Nef from HIV-1 subtype B.
Protocal of our sequence collection
For HIV-1 subtype B, one sequence per patient was extracted from HIV Los Alamos database (www.hiv.lanl.gov/).
We removed misclassified sequences or sequences with hypermutations, stop codons, ambiguous nucleotides, which were described in our article [1].
We removed sequences conferred partial or full resistance to any of the Nef inhibitors, RT inhibitors and Nef inhibitors using HIVdb V6.0 .
Visualization
Our sequence dataset of HIV-1 subtype B Nef included 4725 sequences. In the following picture, HXB2 indices of individual proteins are shown on top of the colored bars. A consensus amino acid at each position is shown beneath the colored bar. Natural variations are shown below the consensus amino acids; proportions (%) are colored red if they were more than 5%; blue otherwise.
HIV-1 protein interaction patterns.
Please cite our article:
Guangdi Li, Supinya Piampongsant, Nuno Rodrigues Faria, Arnout Voet, Andrea-Clemencia Pineda-Peña, Ricardo Khouri, Philippe Lemey, Anne-Mieke Vandamme, Kristof Theys. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18, doi:10.1186/s12977-015-0148-6 (2015). [PDF] [PubMed Link]