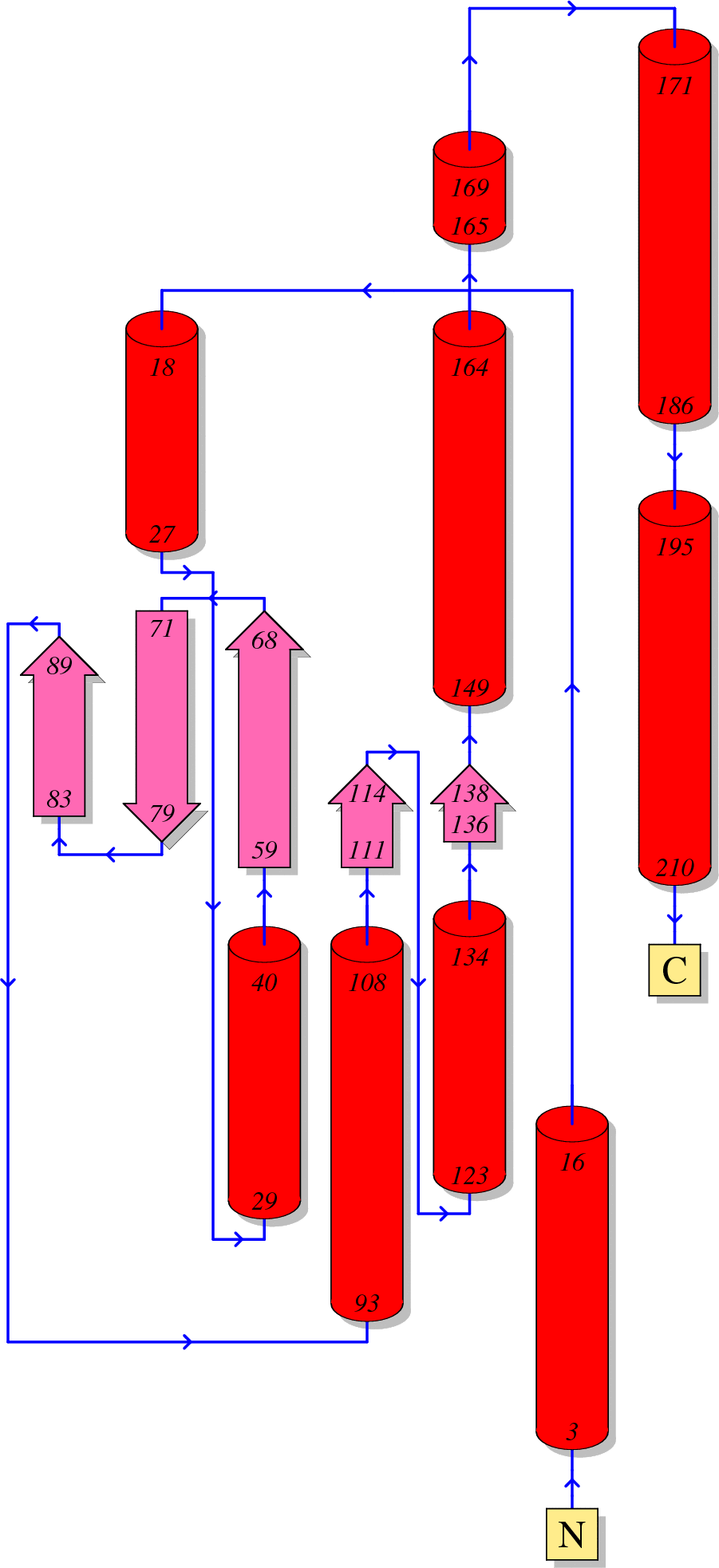
Function of HIV Integrase
Integrase is the third enzyme encoded by the pol gene.
After the nuclear import of pre-integration complex (PIC), viral Integrase catalyzes two major reactions (3’-processing and strand transfer reactions) to insert the linear, double-stranded viral DNA into human chromosomes.
In the mature viral particles, Integrase is cleaved from the GagPol polyprotein by viral Protease. Moreover, reverse transcriptase binds with integrase to prevent the catalytic activity of Integrase before viral integration [1].
As part of reverse transcriptase complex, Integrase also plays a role during reverse transcription [2].
Reference
Zhu K, Dobard C, Chow SA: Requirement for integrase during reverse transcription of human immunodeficiency virus type 1 and the effect of cysteine mutations of integrase on its interactions with reverse transcriptase. J Virol 2004, 78:5045-5055.(Download Article)
Xue B, Mizianty MJ, Kurgan L, Uversky VN: Protein intrinsic disorder as a flexible armor and a weapon of HIV-1. Cell Mol Life Sci 2012, 69:1211-1259.(Download Article)
Sequence
(1) Reference sequence for HIV-1 Integrase
1 10 20 30 40 50
| | | | | |
FLDGIDKAQD EHEKYHSNWR AMASDFNLPP VVAKEIVASC DKCQLKGEAM
51 60 70 80 90 100
| | | | | |
HGQVDCSPGI WQLDCTHLEG KVILVAVHVA SGYIEAEVIP AETGQETAYF
101 110 120 130 140 150
| | | | | |
LLKLAGRWPV KTIHTDNGSN FTGATVRAAC WWAGIKQEFG IPYNPQSQGV
151 160 170 180 190 200
| | | | | |
VESMNKELKK IIGQVRDQAE HLKTAVQMAV FIHNFKRKGG IGGYSAGERI
201 210 220 230 240 250
| | | | | |
VDIIATDIQT KELQKQITKI QNFRVYYRDS RNPLWKGPAK LLWKGEGAVV
251 260 270 280 288
| | | | |
IQDNSDIKVV PRRKAKIIRD YGKQMAGDDC VASRQDED (2) Reference sequence for HIV-2 and SIV Integrase
1 10 20 30 40 50
| | | | | |
FLEKIEPAQE EHDKYHSNVK ELVFKFGLPR IVARQIVDTC DKCHQKGEAI
51 60 70 80 90 100
| | | | | |
HGQANSDLGT WQMDCTHLEG KIIIVAVHVA SGFIEAEVIP QETGRQTALF
101 110 120 130 140 150
| | | | | |
LLKLAGRWPI THLHTDNGAN FASQEVKMVA WWAGIEHTFG VPYNPQSQGV
151 160 170 180 190 200
| | | | | |
VEAMNHHLKN QIDRIREQAN SVETIVLMAV HCMNFKRRGG IGDMTPAERL
201 210 220 230 240 250
| | | | | |
INMITTEQEI QFQQSKNSKF KNFRVYYREG RDQLWKGPGE LLWKGEGAVI
251 260 270 280 290
| | | | |
LKVGTDIKVV PRRKAKIIKD YGGGKEVDSS SHMEDTGEAR EVA (3) Coloring scheme for above amino acids
Amino acids with hydrophobic side chains (normally buried inside the protein core):
A - Ala - Alanine
I - Ile - Isoleucine
L - Leu - Leucine
M - Met - Methionine
V - Val - Valine
Amino acids with polar uncharged side chains (may participate in hydrogen bonds):
N - Asn - Asparagine
Q - Gln - Glutamine
S - Ser - Serine
T - Thr - Threonine
Amino acids with positive charged side chains:
H - His - Histidine
K - Lys - Lysine
R - Arg - Arginine
Amino acids with negative charged side chains:
D - Asp - Aspartic acid
E - Glu - Glutamic acid
Amino acids with aromatic side chains:
F - Phe - Phenylalanine
Y - Tyr - Tyrosine
W - Trp - Tryptophan
Cysteine: C - Cys - Cysteine
Glycine: G - Gly - Glycine
Proline: P - Pro - Proline
Amino acid variations at HIV-1 Integrase
Here, we visualize the prevalence of amino acid variations at the HIV-1 Integrase from HIV-1 subtype B.
Protocal of our sequence collection
For HIV-1 subtype B, one sequence per patient was extracted from HIV Los Alamos database (www.hiv.lanl.gov/).
We removed misclassified sequences or sequences with hypermutations, stop codons, ambiguous nucleotides, which were described in our article [1].
We removed sequences conferred partial or full resistance to any of the Integrase inhibitors, RT inhibitors and integrase inhibitors using HIVdb V6.0 .
Visualization
Our sequence dataset of HIV-1 subtype B Integrase included 4725 sequences. In the following picture, HXB2 indices of individual proteins are shown on top of the colored bars. A consensus amino acid at each position is shown beneath the colored bar. Natural variations are shown below the consensus amino acids; proportions (%) are colored red if they were more than 5%; blue otherwise.
HIV-1 protein interaction patterns.
Please cite our article:
Guangdi Li, Supinya Piampongsant, Nuno Rodrigues Faria, Arnout Voet, Andrea-Clemencia Pineda-Peña, Ricardo Khouri, Philippe Lemey, Anne-Mieke Vandamme, Kristof Theys. An integrated map of HIV genome-wide variation from a population perspective. Retrovirology 12, 18, doi:10.1186/s12977-015-0148-6 (2015). [PDF] [PubMed Link]